Commits on Source (11)
Showing
- .gitignore 1 addition, 0 deletions.gitignore
- .pre-commit-config.yaml 22 additions, 0 deletions.pre-commit-config.yaml
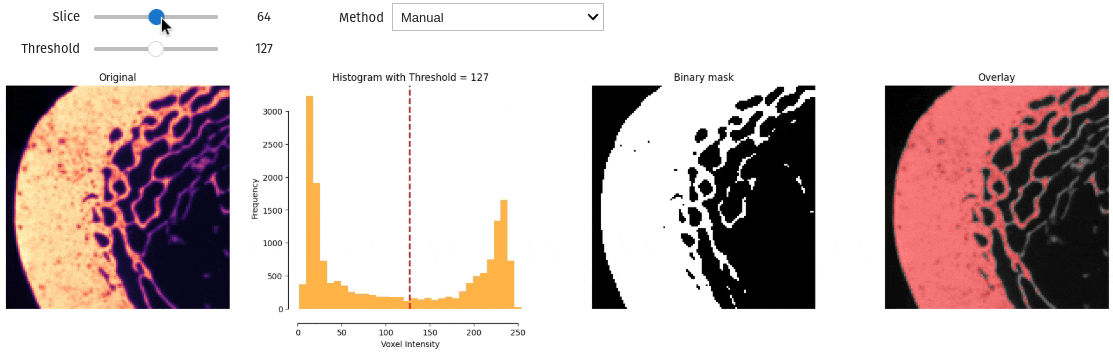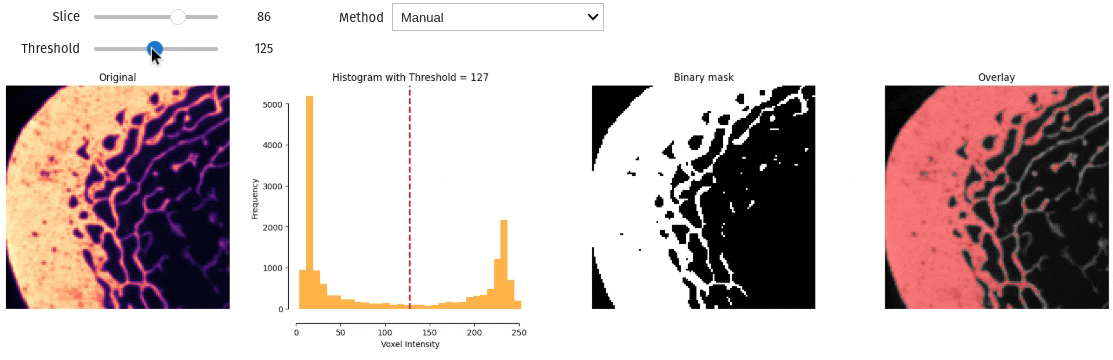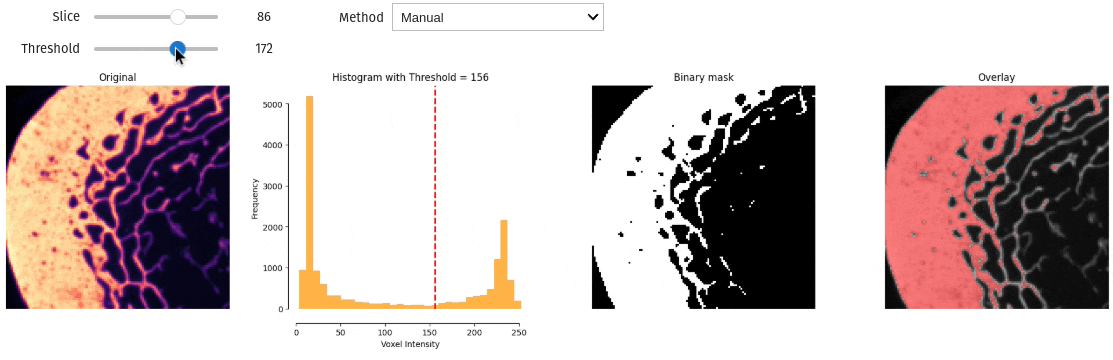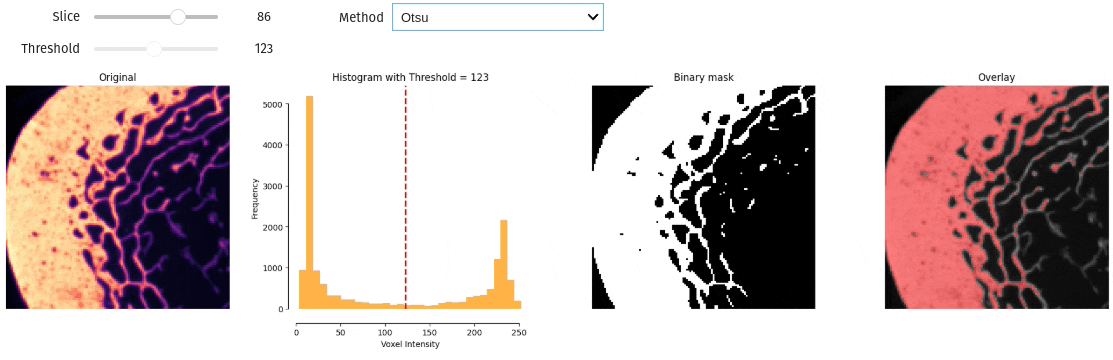
- docs/assets/screenshots/interactive_thresholding.gif 0 additions, 0 deletionsdocs/assets/screenshots/interactive_thresholding.gif
- docs/assets/screenshots/pygel3d_visualization.png 0 additions, 0 deletionsdocs/assets/screenshots/pygel3d_visualization.png
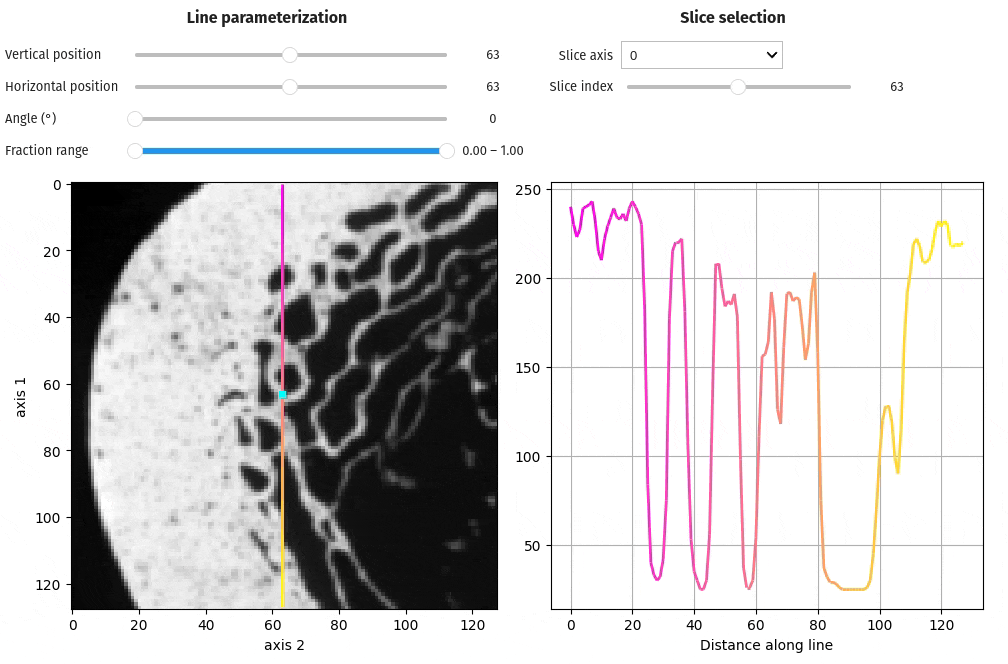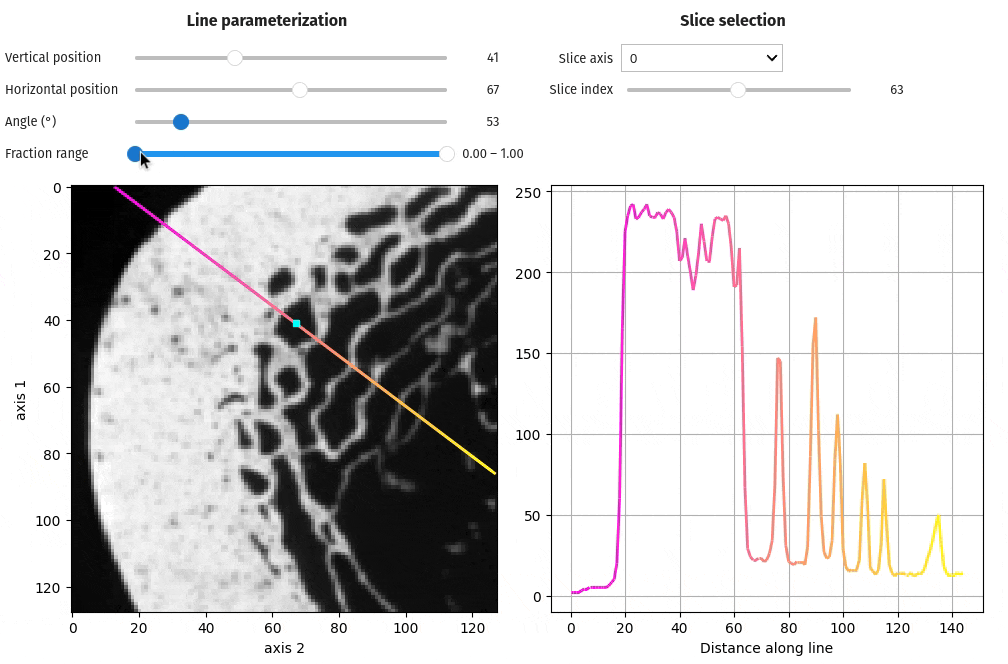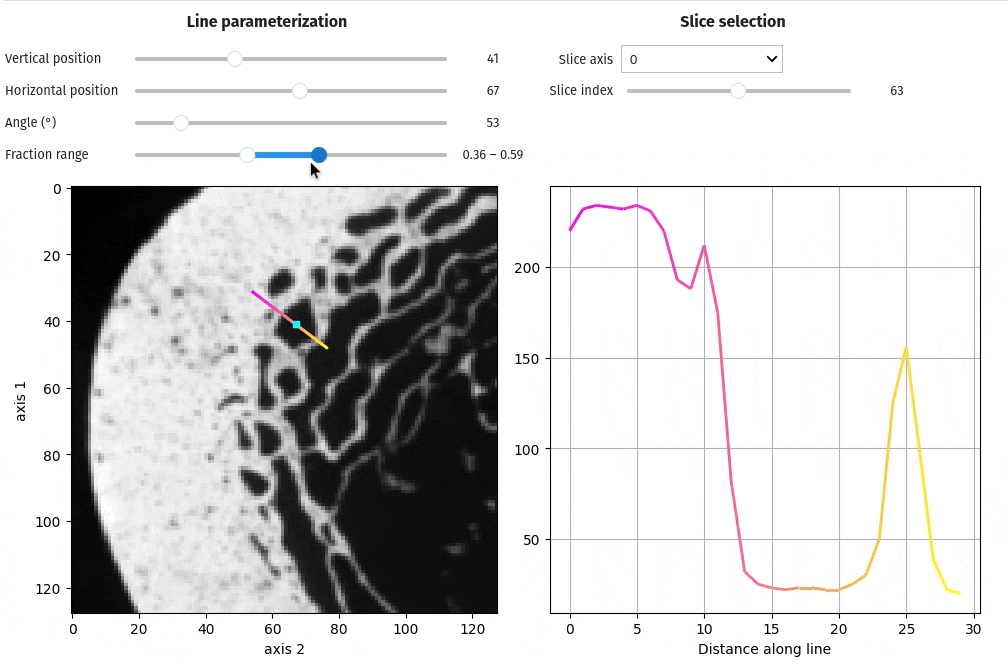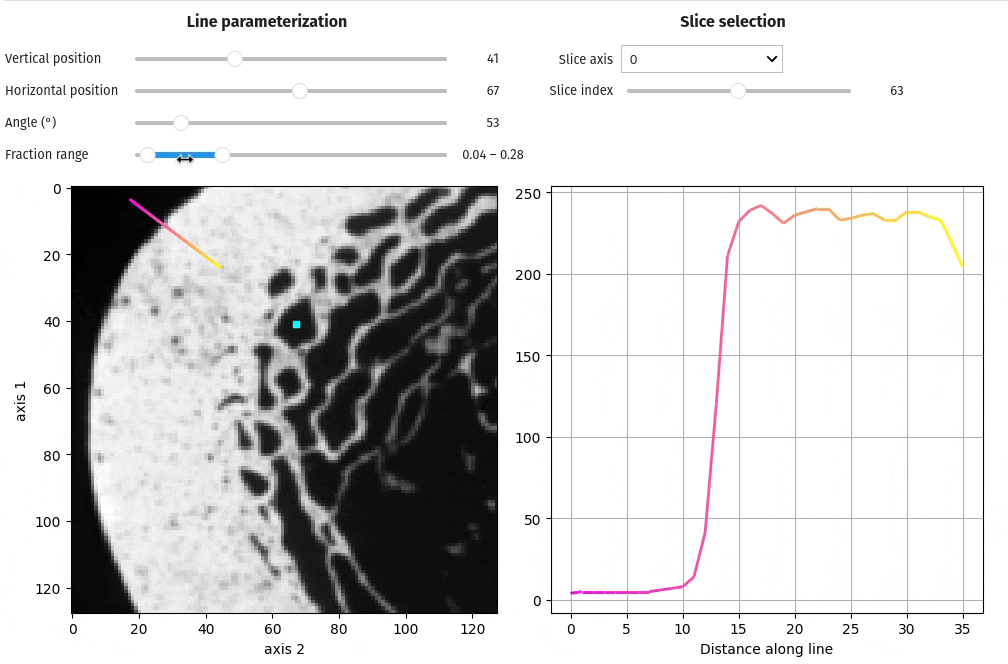
- docs/assets/screenshots/viz-line_profile.gif 0 additions, 0 deletionsdocs/assets/screenshots/viz-line_profile.gif
- docs/doc/cli/cli.md 1 addition, 1 deletiondocs/doc/cli/cli.md
- docs/doc/data_handling/io.md 2 additions, 2 deletionsdocs/doc/data_handling/io.md
- docs/doc/gui/gui.md 1 addition, 1 deletiondocs/doc/gui/gui.md
- docs/doc/releases/releases.md 1 addition, 1 deletiondocs/doc/releases/releases.md
- docs/doc/visualization/viz.md 2 additions, 0 deletionsdocs/doc/visualization/viz.md
- pyproject.toml 92 additions, 0 deletionspyproject.toml
- qim3d/__init__.py 6 additions, 4 deletionsqim3d/__init__.py
- qim3d/cli/__init__.py 68 additions, 74 deletionsqim3d/cli/__init__.py
- qim3d/detection/__init__.py 1 addition, 1 deletionqim3d/detection/__init__.py
- qim3d/detection/_common_detection_methods.py 12 additions, 9 deletionsqim3d/detection/_common_detection_methods.py
- qim3d/examples/__init__.py 5 additions, 4 deletionsqim3d/examples/__init__.py
- qim3d/features/__init__.py 1 addition, 1 deletionqim3d/features/__init__.py
- qim3d/features/_common_features_methods.py 38 additions, 64 deletionsqim3d/features/_common_features_methods.py
- qim3d/filters/__init__.py 1 addition, 1 deletionqim3d/filters/__init__.py
- qim3d/filters/_common_filter_methods.py 106 additions, 65 deletionsqim3d/filters/_common_filter_methods.py
.pre-commit-config.yaml
0 → 100644
2.57 MiB
197 KiB
docs/assets/screenshots/viz-line_profile.gif
0 → 100644
4.74 MiB
pyproject.toml
0 → 100644